Running simulations
To run simulations with SuSMoST one needs
Python interpreter with installed SuSMoST Python packages
An adsorption model
A Python script with simulation code
SuSMoST Python packages installation is described in Installing SuSMoST. Adsorption models are essentially directories with several files. They can be downloaded from the Downloads page, or found in a models directory of SuSMoST installation. Most of models included in SuSMoST contains example.py scripts with exemplar simulation code.
The simplest simulation script (NH3/V3C2 model)
For example lets try to do simulation for the NH3/V3C2 model. First of all lets activate SuSMoST’s Python environment:
$ source ve/bin/activate
Then change your current directory to the model’s directory:
$ cd models/NH3-V3C2
Then launch the example.py script:
$ python example.py
If everything has been done right, in some second you will see the output:
2.7487196394264197
This is a natural logarith of the model’s partition function per site. If you are interested in details, take a look at the source of the example.py script.
Along with the simple example.py script the NH3-V3C2 directory also contains more complicated and usefull example-isotherm.py. It prints out surface coverages for a sequence of NH3/V3C2 adsorption complex energies:
-0.3 0.9554971864503797
-0.24 0.8563763324198087
-0.18 0.7513212340924432
-0.12 0.6294038161599375
-0.06 0.5028657967144572
0.0 0.3445965667110924
0.06 0.1522997166803156
0.12 0.028102706082451023
0.18 0.0030560681307699716
0.24000000000000005 0.00030207886450439256
0.3 2.9554309932887306e-05
Here is an isotherm plot from these data:

The simplest simulation script with MPI (CO/Pt(111) model)
Simulations scripts that use parallel tempering algorithm should be launched appropriately. An example of such script is the example.py file in CO/Pt(111) model. To launch this script you need activate Python environment with SuSMoST and set working directory to the model directory, just like in previous example:
$ source ve/bin/activate
$ cd models/CO-Pt111
But you need to launch such scripts via mpiexec command with number of processes that is equal to number of parallel replicas requested by the script. Number of replicas correspond to the lenght of temperatures list that is passed to the :py:func:make_metropolis function. The models/CO-Pt111/example.py script requests four temperatures, hence it should be launched in this way:
$ mpiexec -n 4 python example.py
Successfull simulation should print out statistical digest about simulation. Like this:
Temperature # 0 = 2.0
Parameter # 0 : 'coverage'
Sample size 1000
Sample mean 0.307809
Sample stdDev 0.000183
Autocorrelation function parameters:
First zero time: 30
Exponential time: 10.556 (RMSE=0.038)
Integrated time: 27.107
Effective sample size integrated 36.891
Effective sample size exponential 47.366
Sample mean stdDev 0.000
Parameter # -2 : 'Energy'
Sample size 1000
Sample mean -0.003104
Sample stdDev 0.000000
Autocorrelation function parameters:
First zero time: 24
Exponential time: 8.418 (RMSE=0.054)
Integrated time: 14.961
Effective sample size integrated 66.839
Effective sample size exponential 59.399
Sample mean stdDev 0.000
Correlations between parameters:
1.000 0.865
0.865 1.000
Temperature # 1 = 8.965331703353698
Parameter # 0 : 'coverage'
Sample size 1000
Sample mean 0.306266
Sample stdDev 0.001398
Autocorrelation function parameters:
First zero time: 66
Exponential time: 16.284 (RMSE=0.030)
Integrated time: 23.136
Effective sample size integrated 43.223
Effective sample size exponential 30.704
Sample mean stdDev 0.000
Parameter # -2 : 'Energy'
Sample size 1000
Sample mean -0.003090
Sample stdDev 0.000006
Autocorrelation function parameters:
First zero time: 93
Exponential time: 26.992 (RMSE=0.028)
Integrated time: 35.761
Effective sample size integrated 27.964
Effective sample size exponential 18.524
Sample mean stdDev 0.000
Correlations between parameters:
1.000 -0.694
-0.694 1.000
Temperature # 2 = 10.695830923113437
Parameter # 0 : 'coverage'
Sample size 1000
Sample mean 0.304536
Sample stdDev 0.002205
Autocorrelation function parameters:
First zero time: 49
Exponential time: 11.973 (RMSE=0.040)
Integrated time: 12.767
Effective sample size integrated 78.325
Effective sample size exponential 41.762
Sample mean stdDev 0.000
Parameter # -2 : 'Energy'
Sample size 1000
Sample mean -0.003076
Sample stdDev 0.000010
Autocorrelation function parameters:
First zero time: 101
Exponential time: 24.256 (RMSE=0.027)
Integrated time: 28.377
Effective sample size integrated 35.240
Effective sample size exponential 20.613
Sample mean stdDev 0.000
Correlations between parameters:
1.000 -0.615
-0.615 1.000
Temperature # 3 = 12.0
Parameter # 0 : 'coverage'
Sample size 1000
Sample mean 0.302917
Sample stdDev 0.002512
Autocorrelation function parameters:
First zero time: 42
Exponential time: 10.671 (RMSE=0.042)
Integrated time: 12.250
Effective sample size integrated 81.634
Effective sample size exponential 46.855
Sample mean stdDev 0.000
Parameter # -2 : 'Energy'
Sample size 1000
Sample mean -0.003066
Sample stdDev 0.000012
Autocorrelation function parameters:
First zero time: 63
Exponential time: 17.843 (RMSE=0.036)
Integrated time: 21.985
Effective sample size integrated 45.485
Effective sample size exponential 28.022
Sample mean stdDev 0.000
Correlations between parameters:
1.000 -0.667
-0.667 1.000
Recommended temperatures: [2.0, 8.119878894794397, 10.375217380314396, 12.0]
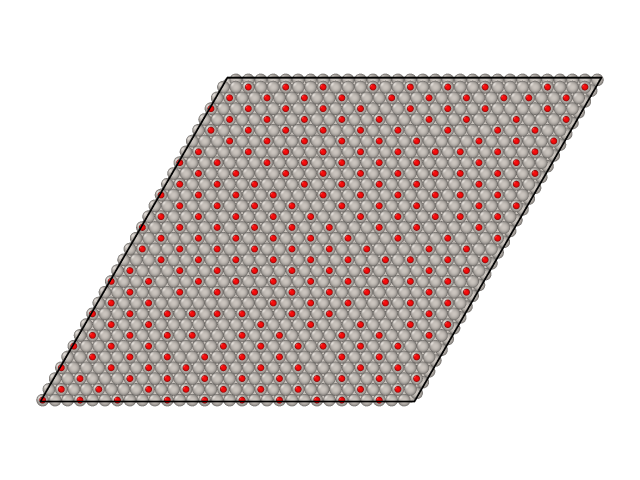
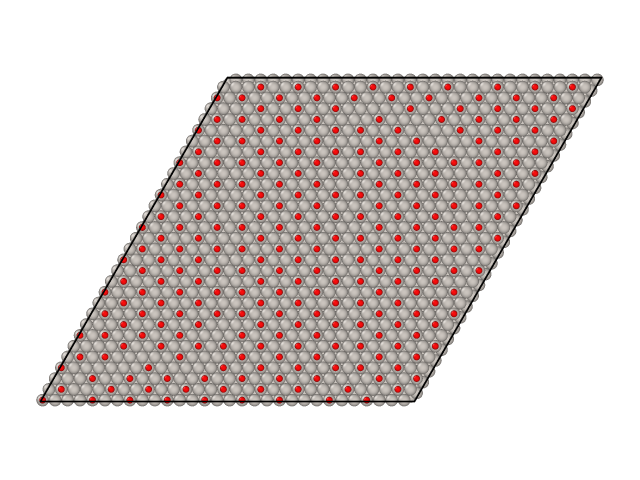
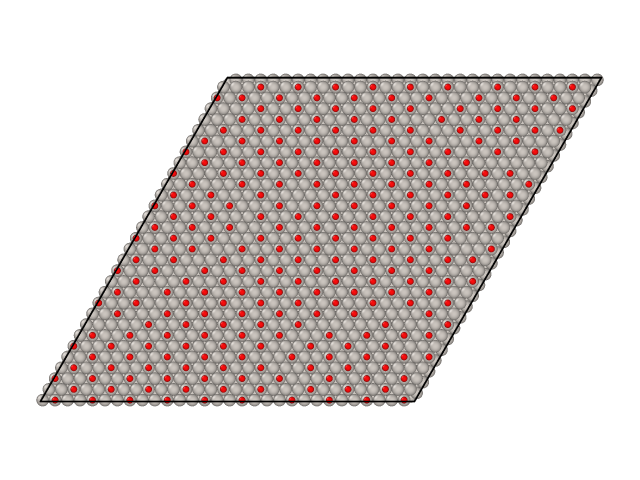
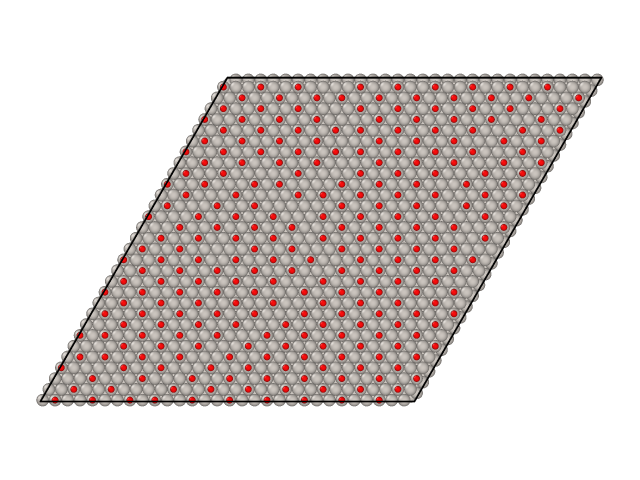
Parallel tempering simulations also generate surface samples as XYZ files. By default these samples are stored in files named traj_T.xyz, where T is the temperature of the sample. Here are samples from CO/Pt(111) simulations we ran above.
|
|
|
|