Tutorial - On manybody interaction
In this tutorial we provide an example of applying the TPD spectra to simulate Langmuir adsorption model with interaction between nearest neighbors and manybody interaction. In this tutorial we use SUrface Science MOdeling and Simulation Toolkit (SuSMoST).
For the simulation of three-particle interactions the calculation of W (the tensor interaction) has changed. The arrangement of molecules for a triangular lattice is determined as follows
cell = Cell([[a,0.,0.], [0.5*a, 0.5*sqrt(3.)*a, 0.], [0.,0.,1.] ] )
here the lattice constant a specifies the location of the nodes.
We see from the Fig.1 that three-particle interactions is ijl and ikl.
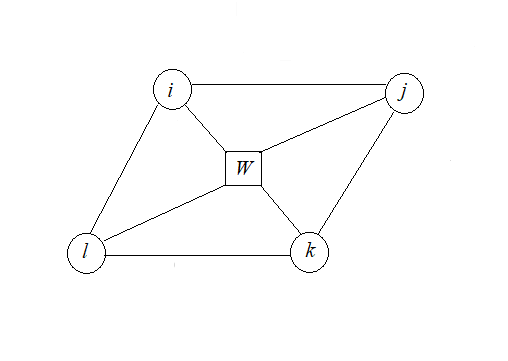
Fig. 1. The arrangement of molecules for a triangular lattice.
Finally, it is necessary to check that all lattice cells are occupied (busy_state_idx), then W is calculated as follows
W[1][busy_state_idx,:,busy_state_idx, busy_state_idx] *= exp(-mol_int3*beta)
W[1][busy_state_idx,busy_state_idx,:, busy_state_idx] *= exp(-mol_int3*beta)
Results
In the Fig.2, Fig.3 and Fig.4 the TPD spectra to simulate Langmuir adsorption model with different interactions: interaction between nearest neighbors, manybody interaction (in this models we use a triangular lattice and N=6 - width of the semi-infinite system (infinite in one direction and finite in perpendicular one), mol_int = 4 is an interaction energy between nearest neighbors, mol_int3 = 3 is an manybody interaction energy) are shown.
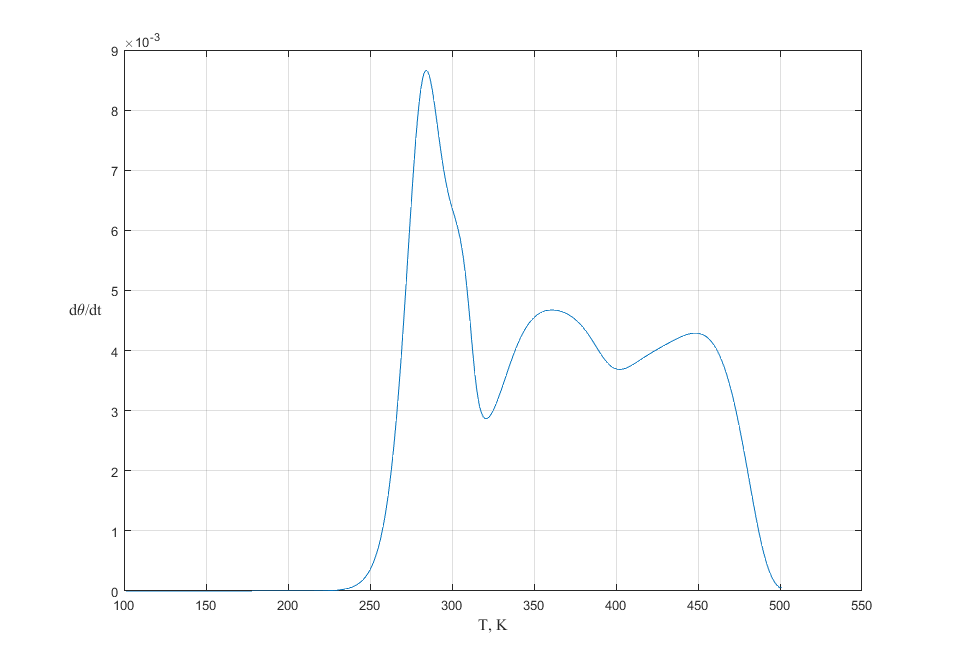
Fig. 2. TPD spectra with heating rates of \(\beta = 20\) K/s. The initial coverage is 1.00. Langmuir adsorption model with interaction between nearest neighbors.
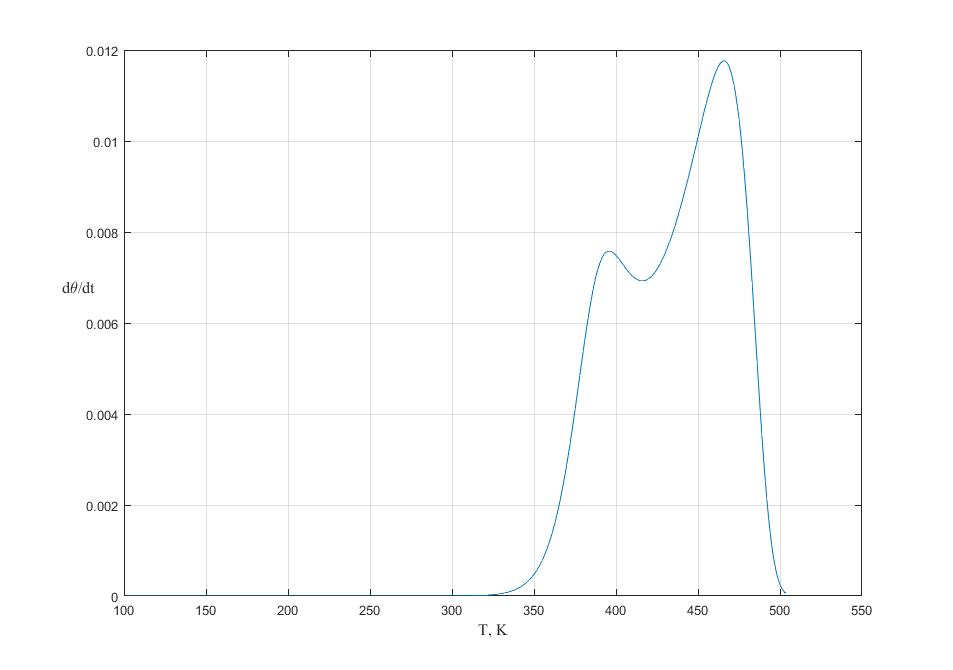
Fig. 3. TPD spectra with heating rates of \(\beta = 20\) K/s. The initial coverage is 1.00. Langmuir adsorption model with manybody interaction.
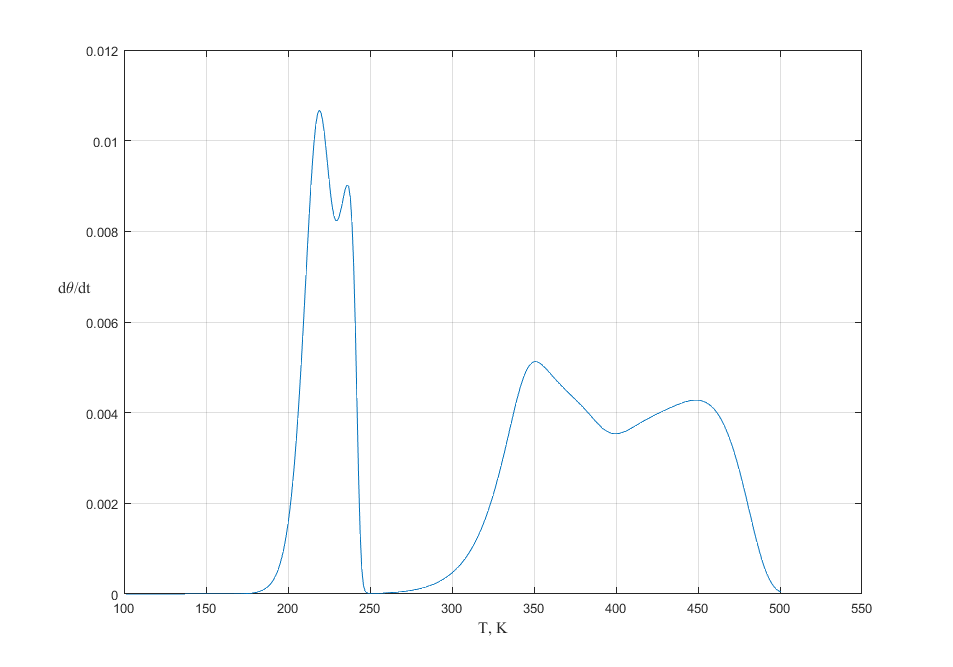
Fig. 4. TPD spectra with heating rates of \(\beta = 20\) K/s. The initial coverage is 1.00. Langmuir adsorption model with interaction between nearest neighbors and manybody interaction.